High-Throughput Phenotyping on a Budget? Yes, Peas!
Researchers Develop Greenhouse-Based Platform to Assess Aphanomyces Root Rot Resistance in Field Pea

Rapid technological advances are ushering in an era of accessibility and affordability for plant phenotyping.
From digital and thermal cameras to 3‐D and fusion approaches, different types of imaging are allowing plant breeders and researchers to screen plant phenotypes quickly and efficiently, saving hundreds of hours and accelerating progress.
In a recent paper in The Plant Phenome Journal, researchers developed and tested a greenhouse‐based high‐throughput phenotyping platform using a digital camera that allowed them to identify and then genetically dissect resistance to Aphanomyces root rot in field pea.
Rapid technological advances are ushering in an era of accessibility and affordability for plant phenotyping.
From digital and thermal cameras to 3‐D and fusion approaches, different types of imaging are allowing plant breeders and researchers to screen plant phenotypes quickly and efficiently, saving hundreds of hours and accelerating progress.
In a recent paper in The Plant Phenome Journal, researchers developed and tested a greenhouse‐based high‐throughput phenotyping platform using a digital camera that allowed them to identify and then genetically dissect resistance to Aphanomyces root rot in field pea.
Rapid technological advances inside and outside the field of plant breeding are ushering in an era of accessibility and affordability for plant phenotyping. From digital and thermal cameras to 3‐D and fusion approaches, different types of imaging are allowing plant breeders and researchers to screen plant phenotypes quickly and efficiently, saving hundreds of hours and accelerating progress.
In plant breeding, the most difficult traits to select for are called quantitative traits because they arise from the effects of hundreds or even thousands of genes. This can make the breeding process very challenging when trying to make improvements to one of these traits; precision techniques like CRISPR gene editing may not be effective due to the complex genetics at play. In addition, it’s also possible that researchers have yet to develop the capacity to have a fuller understanding of missing heritability and dissect the genes into their major effects. This is where the emerging field of plant phenomics steps in.
Valerio Hoyos‐Villegas, an assistant professor and plant breeder at McGill University, published work 10 years ago in Crop Science on how inexpensive hand‐held digital cameras can successfully assess soybean growth and yield. Since then, the CSSA and ASA member says this kind of work has come a long way, and now more tools are available that provide much more detailed information at a significantly lower cost.
“Today, if you have someone with knowledge of electronic engineering and a bit of coding, you can even build your own proprietary or custom tools to measure whatever it is you are interested in phenotyping,” he says. “This is the idea that we can pursue tools and procedures that we can use in order to dissect quantitative trait variation or analyze plant characteristics from alternative perspectives. This means taking measurements more effectively and efficiently than the human eye or even beyond what humans can observe.”
Getting to the Root of Aphanomyces Root Rot
CSSA member Nonoy Bandillo is an assistant professor and plant breeder at North Dakota State University who is exploring techniques to develop disease‐resistant trait packages for pulse crops, including field pea. In a recent paper in The Plant Phenome Journal (https://doi.org/10.1002/ppj2.20063), he and collaborators Abdullah Al Bari, Julie Pasche, and Paulo Flores (CSSA and ASA members) developed and tested a greenhouse‐based high‐throughput phenotyping platform using a digital camera that allowed them to identify and then genetically dissect resistance to Aphanomyces root rot in field pea.
“In this work, farmers need early maturing high‐yielding cultivars with superior quality attributes and improved disease resistance, and in this case, we are focused on Aphanomyces root rot,” Bandillo explains. “The evaluation of root rot on plants in the greenhouse requires a trained expert and is extremely challenging, time consuming, and prone to human error because a breeding program can have thousands of lines. So we wanted to develop a tool that would speed up that scoring process where plants are evaluated for resistance to root rot.”
Aphanomyces root rot is caused by Aphanomyces euteiches. By infecting plant roots, it makes them unable to take up water or nutrients, slowly killing the plant. Symptoms of the disease also begin to show in the leaves of the plant, which turn yellow as they die, and this is what Bandillo and colleagues hoped to take advantage of. Their goal was a tool that balanced efficacy and cost that could be useful to the program.
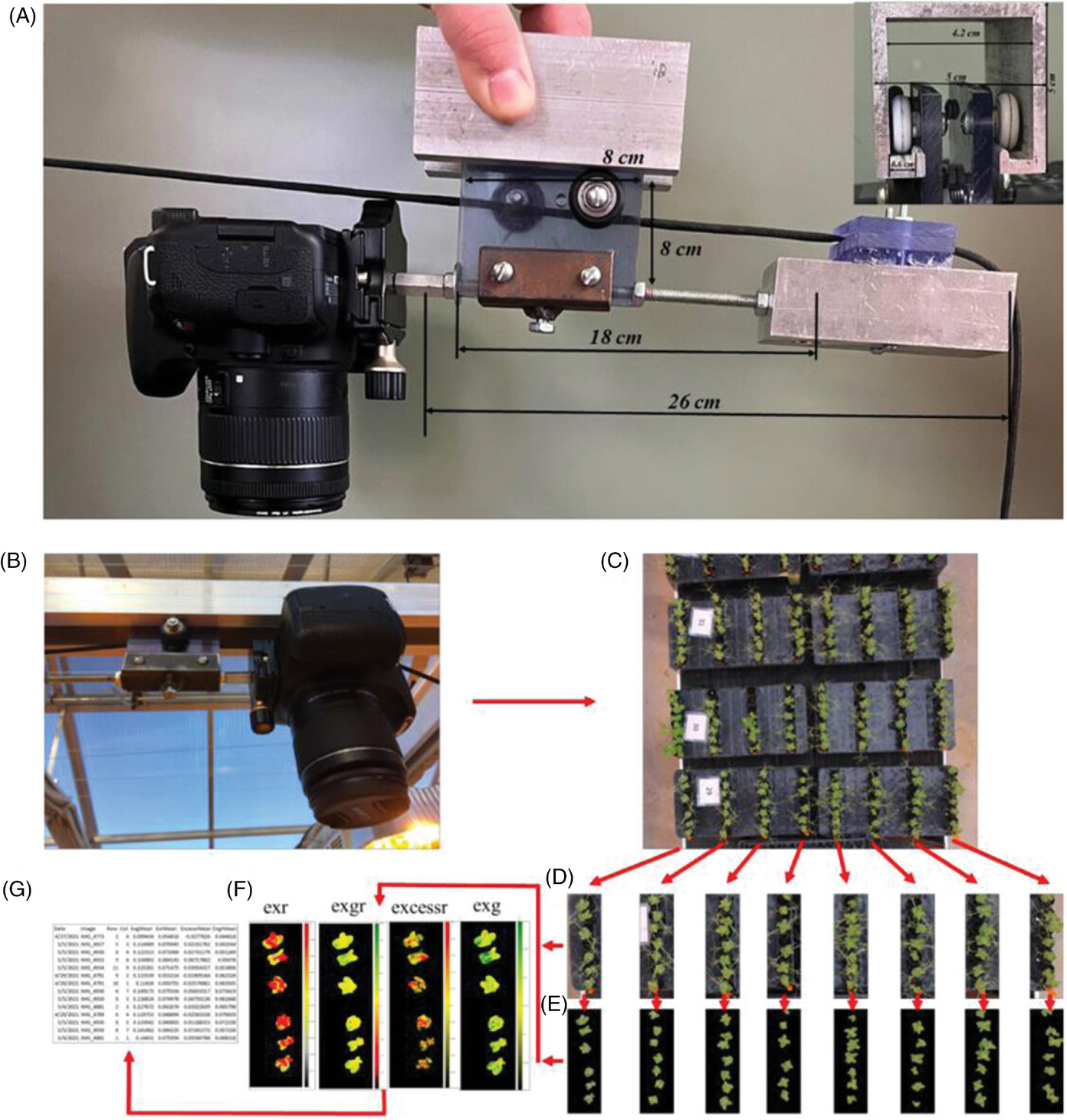
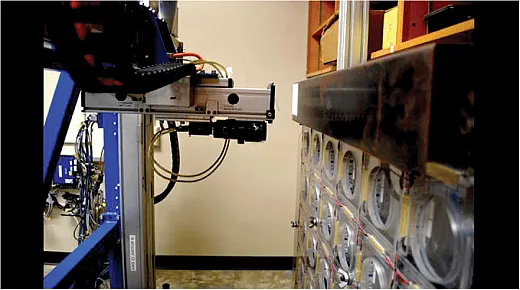
They developed a greenhouse‐based high‐throughput phenotyping platform that utilizes a mounted digital camera and moves along rails above the greenhouse benches (Figure 1). The camera captures images of the visual foliar symptoms of root rot in the plant canopy. The platform is able to image more than 2,000 plants quickly.

To determine the likelihood that a plant is showing symptoms of root rot, the numerous images are then translated into numerical values in RGB indices by analyzing the percentage of pixels in an image that are yellow or brown versus green, for example. Individual plant scores are then averaged for a particular line in the breeding study. In conventional scoring, a human expert observes a plant, approximates how much is yellow or brown versus green, and assigns a score (Figure 2).
“We then wanted to know how biologically meaningful the data was from a breeding perspective, and we found that the broad‐sense heritability was higher at 86% for our RGB‐derived indices versus 59% for conventional visual scores,” Bandillo says. “Our analysis also showed that the RGB‐derived indices achieved about 70% accuracy in predicting disease severity, which was higher than the conventional scoring method.”
Next, the researchers used DNA sequencing and genome‐wide association mapping to search for variants in specific parts of DNA—called single nucleotide polymorphisms—that account for root rot resistance. This helps plant breeders know if a gene or set of genes are a good target for gene editing. With more work in the greenhouse, followed by field assessments, the researchers’ findings can help advance cultivar development for root rot resistance.
An Affordable, Accessible, and Powerful Approach
They performed genome‐wide association mapping with both conventional and the high‐throughput scoring and found consistently higher numbers of single nucleotide polymorphisms with the high‐throughput method. Their high‐throughput work identified previously mapped genes that are involved in conferring immunity against the root rot as well as some novel quantitative trait loci, which are regions of DNA associated with a complex quantitative trait. These loci will inspire future work.
“This means the high‐throughput method has more power for narrowing down candidate genes than the conventional method,” Bandillo says. “We found this technique to be low‐cost and accurate enough to be helpful in our research. Because the heritability is higher, we now have more power to detect DNA associated with our disease, which was not being detected before. This is the first study that we know of that reported on the implementation of a greenhouse‐based phenotyping platform to assess disease resistance in peas.”
While effective, the technique does have some limitations, the scientists note. One being that the camera only captures images of the leaves themselves, not the roots where the disease is present. This means that tolerant lines, where plants are infected with root rot but have no visible foliar symptoms, can impact the classifying of resistant and susceptible genotypes. However, the study found a high accuracy of nearly 70% for predicting disease severity and further greenhouse and field validations can overcome these limitations, Bandillo says. The platform is most useful for early stages of testing where breeders are screening thousands of lines to eliminate unfavorable genotypes (e.g., susceptible plants).
The value of using high‐throughput phenotyping technology is often twofold: it is a cheaper and more efficient use of human resources, and it can capture better data. A breeding study may be evaluating 300 plant lines, for example, with seven plants per line. At more than 2,000 plants total, that can take human scorers about a week or two. Once an affordable high‐throughput phenotyping tool is developed, it can have data ready in a few hours.
“When I talk about this work and people hear me describe a new tool, some think it’s expensive,” Bandillo says. “In this case, it’s really not. You just need a camera to capture images that can be purchased at any electronics store. It makes this affordable and accessible. I want to solve what I call this phenotyping bottleneck problem that has been here for quite a long time. It is personal to me because I’m a plant breeder and these technologies allow me to allocate more of my effort to doing things that can advance this work.”

While the use of a digital camera for high‐throughput phenotyping is one of the cheapest options, there are many others that have become relatively less expensive over the last several years. Hoyos‐Villegas points to multispectral imaging from cameras and line sensors, as well as thermal imaging, which has fallen in price about tenfold in the last seven years thanks to advances in the technology and now comes in compact versions that can be attached to a phone or drone. While they can still be expensive, 3‐D cameras have also become more affordable. More sophisticated options include Lidar (Light Detection and Ranging), hyperspectral, and full pulse florescence imaging.
“The lower costs not only enable people to utilize these sensors, but also take sensor fusion approaches,” Hoyos‐Villegas explains. “This is where scientists bring together different sensors to capture or characterize a plant trait that is not visible to a single sensor, much less the human eye. It is only reconstructable via the accumulation of data from two different sensors. One example is the combination of both 3‐D and thermal imagining.”
“When I talk about this work and people hear me describe a new tool, some think it’s expensive. In this case, it's really not."
Leveling the Playing Field
He adds that more access to analytics tools and big data computing clusters is also fueling progress because more people now have the ability to analyze and understand the large amounts of data they can capture. In the recent past, this work may have been more limited to professional phenotyping facilities at large universities and the private sector.

Hoyos‐Villegas also says these decreasing costs have helped level the playing field in some ways when it comes to the size, scope, and budgets of breeding programs. When technologies are more expensive, only larger university programs and the private sector working on mainstream, well‐supported crops can afford them, leaving smaller programs focused on specialty commodities behind. When technologies become more affordable and are developed for use in phenotyping, smaller programs can also adopt the techniques.
Hoyos‐Villegas imagines the opportunity to create a kind of federated system where breeders partner to share data they collect using new technologies. Smaller breeding programs working on specialty crops can greatly benefit from this approach by pooling data to train better models and make more accurate predictions than their own data would allow.
“This is all still coming of age, and there is a lot of buzz around it,” Hoyos‐Villegas says. “While I believe plant breeders have yet to see the first breeding program and plant variety developed fully using these techniques, researchers and industry are beginning to leverage phenomics in new ways. This is really the next wave.”
A key to making this work successful is its interdisciplinary nature. This work is at the intersection of plant science and engineering and can include plant breeders and pathologists and agricultural, chemical, electrical, and materials engineers. Hoyos‐Villegas sees this interdisciplinary work facilitating the practice of plant breeding to be more retrospective.
“As breeding programs progress, we often miss the opportunity to look back and see what we can learn to improve our work,” he says. “But these other disciplines are helping create new knowledge that can be used in a breeding program. We just need a way to bring everyone together, and I think phenomics is one way.”
DIG DEEPER
View the research mentioned in this article:
Bari, Md.A.Al., Fonseka, D., Stenger, J., Zitnick‐Anderson, K., Atanda, S.A., Morales, M., … & Bandillo, N. (2023). A greenhouse‐based high‐throughput phenotyping platform for identification and genetic dissection of resistance to Aphanomyces root rot in field pea. The Plant Phenome Journal, 6, e20063. https://doi.org/10.1002/ppj2.20063
Text © . The authors. CC BY-NC-ND 4.0. Except where otherwise noted, images are subject to copyright. Any reuse without express permission from the copyright owner is prohibited.