Unlocking the Secrets of Soil: Exploring the Microbiome and Its Applications—Part 2
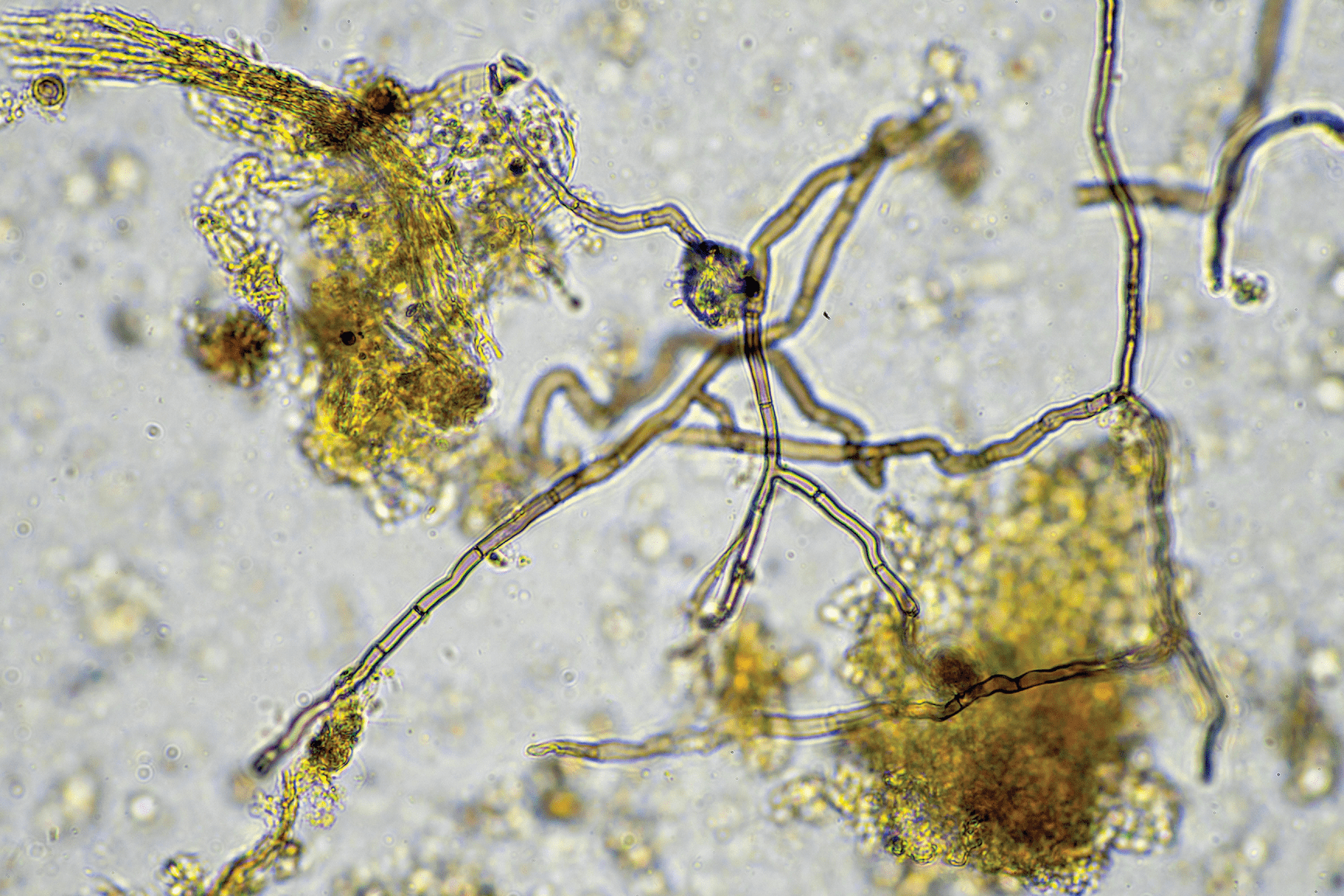
The term “microbiome” refers to a collective assembly of microorganisms within a specific environment, including fungi, bacteria, and viruses, influencing dynamic functions within the system. Microbes play a pivotal role in the production of various food items, wines, and medicinal compounds. Their significance in agriculture and ecology is monumental, which will be explored in this two‐part series. In this second part, we’ll discuss the microbiome’s role in plant adaptation and driving the biogeochemical processes in soil across scales, mathematical and statistical modeling of plant–soil–microbe interactions, and areas of future research.
View the entire series.
Microbiome for Plant Adaptation: A Heritable Trait

Phenotypic plasticity, a crucial trait enabling plants to adapt to diverse environments, involves the modification of their traits, which enhances the plant fitness to the environment. This has led to the evolution of many factors that contribute to phenotypic plasticity, including the alteration in the microbiome of the rhizosphere through root exudates. Several studies have reported that plants can increase their tolerance towards environmental stress like drought, heat, salinity, and other abiotic and biotic stresses by changing the microbiome at the rhizosphere level (Pascale et al., 2020; Zhang et al., 2019; Castrillo et al., 2017).
By modulating the composition and diversity of the microbiome, plants can improve the uptake of nutrients and water and harbor high mycorrhizal activity; thus, capturing the resources from distant levels, which can help with survival of different environmental stresses. Microbes are also considered the second crop genome, regulating crop dynamics and functioning. Advancements in whole‐genome sequencing and modern genomic technologies have provided insights into beneficial plant–microbe interactions in soil, offering potential crop improvement strategies.
It has been reported that plants influence the rhizosphere microbiome, showing unique microbial composition and diversity. For example, drought‐tolerant crops harbor Actinobacteria, which helps combat plant drought stress (Naylor et al., 2017). Genome‐wide association studies in Arabidopsis and other plants have revealed the strong association between host and microbial genome, revealing the heritable interactions that can be a target for future crop breeding and improvement programs. Similarly, investigations into the gut microbiomes of swine, baboons, and humans revealed high heritability of microbiomes (Grieneisen et al., 2021; Bergamaschi et al., 2020). Deng et al (2021) have reported that plant loci control rhizosphere microbiome and cause genotype‐specific microbiome composition. This has led to the development of microbiome heritability concepts.
Microbiome heritability has been defined as a deterministic component of community assembly caused by the host genetics, which can be either deterministic or stochastic over span of time and space (Wagner, 2021). The size of the microbial community has been identified as a crucial determinant of influencing whether the host phenotypes exhibit deterministic or stochastic effects, thereby significantly impacting microbiome heritability (Bisschop et al., 2022; Gao et al., 2020). Similarly, the barley domestication has harbored the microbiota positively by developing resistance against pathogens and increased nutrient and water uptake over the course of time (Bulgarelli et al., 2015). Thus, study of plant–microbe interactions and evaluating microbiome heritability will have a huge positive impact on crop productivity and soil health.
Microbiomes Drive Biogeochemical Processes in Soil Across Scales
The soil biogeochemical cycling, particularly processes that mineralize nutrients bound up in carbonaceous biomass such as soil organic matter, is critical to the functioning of productive ecosystems. Microbes play a crucial role in driving these processes in the soil. Recent advancements in molecular tools and methodologies have provided deeper insights into how specific groups of microbes regulate these functions, shedding light on their activities and abundance. Understanding the ecology of these crucial microbes is critical to establish connections among microbial communities, their activities, and mechanisms governing soil biogeochemical cycling. This comprehension is vital not only for elucidating nutrient‐cycling dynamics but also for assessing the impact of human activities, such as soil, water, and air pollution, on the soil’s capacity to store and sequester atmospheric carbon.
Elucidating the nutrient‐cycling dynamics and ecosystem function requires examining how nutrient inputs shape soil microbial community composition and metabolic capacities. Let’s delve into some of the major microbially mediated biogeochemical processes such as carbon and nitrogen cycling, which are essential for maintaining soil fertility and enhancing ecosystem productivity.
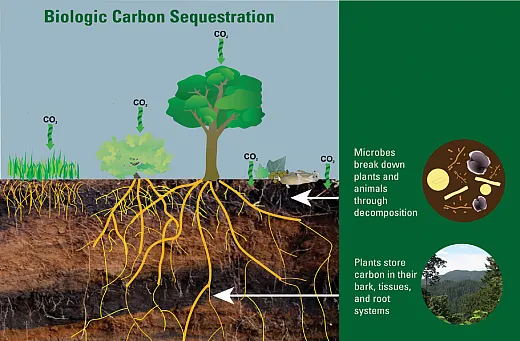
Decomposition of Organic Matter (Carbon Cycling and Sequestration)
Microbes decompose organic matter in the soil through the hydrolysis of macromolecules into lower‐molecular‐weight compounds that can be assimilated to support microbial metabolism. These processes are catalyzed by microbial enzymes, which are affected by microbial activity, composition, nutrient availability, substrate quality, etc. Elucidating carbon cycling principles and mechanisms governing rhizosphere microbiome assembly promises to inform integrated soil management, leveraging plant–microbe interactions to enhance agricultural productivity and sustainability. Furthermore, soil carbon sequestration is notably more reliable and stable in undisturbed environments such as grasslands and forests where root exudates serve as a resource for microbial growth and thus facilitate long‐term sequestration by transitioning of carbon from labile to stable and recalcitrant pools (Panchal et al., 2022).
Increased sequestration is crucial for capturing elevated atmospheric CO2 levels and mitigating global warming. The issue of greenhouse gas emissions has become a major global concern, underscored by initiatives highlighted at events like the United Nations climate change conferences. Estimates, such as those presented by Kell (2012), suggest that a 10% increase in CO2 sequestration in soil could potentially remove up to 20% of CO2 from the atmosphere, which shows the huge potential of microbial‐mediated carbon sequestration for mitigating climate change and global warming.
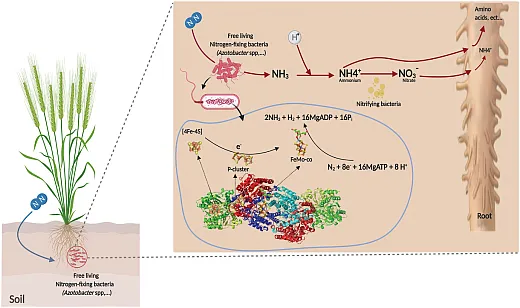
Biological Nitrogen Fixation (Nitrogen Cycling)
Biological nitrogen fixation is a vital process that converts atmospheric nitrogen gas into readily available nitrogen for plant uptake and growth, facilitated by the nitrogenase enzyme system produced by certain soil microbes. Symbiotic nitrogen fixation typically occurs between rhizobia bacteria and legumes while free‐living nitrogen fixers like Azotobacter, Clostridium, and Pseudomonas independently fix nitrogen. Pseudomonas and symbiotic Rhizobium can synergistically enhance nodule formation and nitrogen fixation capacities under some conditions. Further research into microbiome assembly principles and microbial interactions governing nitrogen transformations promises to improve integrated soil management practices that optimize biological nitrogen fixation.
Phosphorus Solubilization (Phosphorus Cycling)
Understanding microbial involvement in rhizosphere phosphorus dynamics require a detailed examination of phosphorus solubilization and its modulation by microbial activities. Various bacteria including actinomycetes, filamentous fungi, and yeasts possess the ability to solubilize both mineral and organic phosphorus forms. However, the same microbes solubilizing phosphorus, or different species, may subsequently immobilize the released phosphorus, limiting plant availability. Mycorrhizal associations are believed to play a crucial role in capturing solubilized phosphorus from the soil solution and storing polyphosphate in fungal structures in soil and roots. This underpins phosphorus–carbon exchange between fungal and plant partners. A holistic understanding of microbial phosphorus solubilization, immobilization, and facilitation of plant phosphorus uptake is needed to manage rhizosphere communities optimizing plant phosphorus acquisition.
Plant–Soil–Microbe Interactions: Mathematical Modeling Approaches
Plant–soil–microbe interactions are fundamental in shaping the rhizosphere and terrestrial ecosystems, encompassing agricultural land, forests, and grasslands, with significant implications for carbon sequestration and food production. These interactions form dynamic networks that can be mutualistic, symbiotic, or antagonistic, influencing nutrient cycling, microbial diversity and activity, soil health, and plant productivity. As global climate change accelerates, ecosystems are rapidly evolving, leading to increased carbon emissions and impacting overall ecology and agriculture. Understanding these components, their interactions, and modeling their future consequences is vital for agricultural and conservation efforts aimed at boosting carbon sequestration and reducing emissions. Modeling these interactions offers a mechanistic understanding of their functioning and helps identify the driving factors.
Plants release exudates containing sugars, amino acids, and other metabolites, serving as energy sources for soil microbiomes. In return, microbes facilitate organic matter decomposition, provide nutrients to plants, fix atmospheric nitrogen, solubilize different nutrients, and enhance soil structure and aggregation. These activities are influenced by the dynamics of each component and the environment, creating a feedback loop where plants and soil influence microbial communities, shaping overall ecosystem dynamics. Let’s see the different approaches of modeling plant–soil–microbe interactions.
We will start with a brief explanation of microbe–microbe interaction.
Advancements in DNA sequencing have revolutionized our understanding of microbe–microbe and plant–microbe interactions. Microbe–microbe interactions can be cooperative, competitive, or neutral based on the resource and environmental stimuli. Assembly theory is regarded as the primary theory for modulation of a microbiome at the community level, which is highly influenced by environmental conditions, resource availability, and interactions with other microbial species (Bai et al., 2022). Microbe–microbe interactions can be studied in general through kinetic, stochastic, and network‐based models. Kinetic models, which are dynamic and deterministic, predict microbial growth, diversity, and activity using differential equations. Stochastic models describe microbial activity in scenarios with lower diversity or asymmetric distributions. The noisy patterns can be removed through Monte Carlo simulations. Network‐based methods help to find the network of interactions among microbial communication for resources and functioning. Overall, integration of theoretical and computational approaches enhances understanding of microbial ecology.
Now, let’s see how we can approach plant–soil–microbe interaction as a whole process.
Modeling interactions among plants, soil, and microbes involves various biological, chemical, and physical processes that connect these systems (Beck et al., 2022). Different approaches have been applied to understand the intricacies of these interactions, ranging from simplistic compartment models to complex mechanistic models. Compartment models approach the understanding of nutrient and energy flow among different components using differential equations, aiding in understanding nutrient cycling and ecosystem dynamics. Dynamic simulation modeling simulates ecosystem dynamics over time, accounting for temporal changes in factors such as soil moisture, nutrients, plant growth, and microbial activity. Metabolic modeling helps in quantifying the stoichiometric matrix of metabolic reactions occurring at the interface of plants and microbes in the soil, elucidating plant–microbe and microbe–microbe interactions (Beck et al., 2022). Agent‐based modeling, including game theory and cellular automata, can be used to study the decision‐making processes by each component in the plant–soil–microbe interface, valuable for optimizing resources in crop production (Almpanis et al., 2019). Stochastic modeling aids in analyzing the diversity and structure of microbial communities at different soil depths and different levels of carbon availability, which overall affect microbial activity and the microbiome (Benucci et al., 2023). In‐Silico‐based modeling of interactions has given the power to derive reference models to simulate different plant–microbe interactions and extend the theoretical framework to the experimental‐based framework.
With the revolution in high‐throughput phenotyping and sequencing, the huge amount of data generated can be analyzed through machine‐learning models. Machine‐learning models perform large‐scale data curation, classification and identification of hidden patterns, and predictions through different parameter estimation and model optimization; thus, deriving hidden relations among the plant–soil–microbial interface.

Microbiome Research is Poised for Exciting Times Ahead
Plant–soil–microbe interactions are pivotal in shaping the rhizosphere and terrestrial ecosystems, spanning from agricultural land to forests and grasslands. These interactions are crucial for carbon sequestration and plant productivity. The intricate relationships among plants, soil, and microbes create a dynamic system that can be mutualistic, symbiotic, or antagonistic, influencing nutrient cycling, microbial diversity and activity, soil health, and plant productivity. With the onset of global climate change, ecosystems are rapidly evolving, leading to increased carbon emissions and affecting overall ecology and agriculture. Understanding these components, their interactions, and modeling their future consequences is essential for agriculture and conservation efforts aimed at increasing carbon sequestration and reducing emissions.
Further, maximizing the capacity and function of soil microbiomes is challenged by both natural and anthropogenic disturbances. Key challenges are discovering novel microbial species and genes governing soil health alongside developing integrated practices that nourish beneficial microbiota while reducing inputs and environmental impacts. Advancing a predictive understanding of microbiome responses to global change is critical for developing biological solutions that harness microbial potential to sustain food production.
To date, although a variety of tools, approaches, and indicators have been developed to characterize microbial composition and functions across scales, ways to integrate these measures to derive a meaningful assessment of ecosystem functioning (e.g., soil health impacts) are often inadequate and still evolving. While individual rhizosphere microorganisms’ roles in biogeochemical processes are increasingly understood, interactions between beneficial groups remain less explored.
Additionally, soil fauna–microbe interactions in the rhizosphere remain relatively understudied. Further insights into microbial community assembly rules, species interactions, and functional genomics promise to inform integrated management leveraging microbial diversity. Thus, to enhance sustainable agriculture, a comprehensive understanding of rhizosphere communities is crucial, particularly regarding their impact on carbon, nitrogen, and phosphorus transformations that supports overall sustainable management of crops. While traditional concepts focus on root–microbe symbioses and microbe–microbe antagonism, there is a need to emphasize network‐level interactions and move beyond binary plant–microbe relationships to incorporate multiscale relationships among plant, microbe, and environments. Exploring assembly principles, species interactions, and microbial trait‐based ecology can help formulate strategies to leverage the soil microbiome for agricultural productivity and sustainability. Additionally, government‐level policy and initiatives on microbiome research and funding hold the future of sustainable agriculture and ecology.

References
Almpanis, A., Corre, C., & Noel, A. (2019). agent based modeling of the rhizobiome with molecular communication and game theory. In Proceedings of the Sixth Annual ACM International Conference on Nanoscale Computing and Communication (pp. 1–7).
Bai, B., Liu, W., Qiu, X., Zhang, J., Zhang, J., & Bai, Y. (2022). The root microbiome: Community assembly and its contributions to plant fitness. Journal of Integrative Plant Biology, 64(2), 230–243.
Beck, A.E., Kleiner, M., & Garrell, A.K. (2022). Elucidating plant‐microbe‐environment interactions through omics‐enabled metabolic modelling using synthetic communities. Frontiers in Plant Science, 13, 910377.
Benucci, G. M. N., Beschoren da Costa, P., Wang, X., & Bonito, G. (2023). Stochastic and deterministic processes shape bioenergy crop microbiomes along a vertical soil niche. Environmental Microbiology, 25(2), 352–366.
Bergamaschi, M., Maltecca, C., Schillebeeckx, C., McNulty, N. P., Schwab, C., Shull, C., … & Tiezzi, F. (2020). Heritability and genome‐wide association of swine gut microbiome features with growth and fatness parameters. Scientific Reports, 10(1), 10134.
Bisschop, K., Kortenbosch, H. H., van Eldijk, T. J., Mallon, C. A., Salles, J. F., Bonte, D., & Etienne, R. S. (2022). Microbiome heritability and its role in adaptation of hosts to novel resources. Frontiers in Microbiology, 13, 703183.
Bulgarelli, D., Garrido‐Oter, R., Münch, P. C., Weiman, A., Dröge, J., Pan, Y., … & Schulze‐Lefert, P. (2015). Structure and function of the bacterial root microbiota in wild and domesticated barley. Cell Host & Microbe, 17(3), 392–403.
Castrillo, G., Teixeira, P. J. P. L., Paredes, S. H., Law, T. F., De Lorenzo, L., Feltcher, M. E., … & Dangl, J. L. (2017). Root microbiota drive direct integration of phosphate stress and immunity. Nature, 543(7646), 513–518.
Deng, S., Caddell, D. F., Xu, G., Dahlen, L., Washington, L., Yang, J., & Coleman‐Derr, D. (2021). Genome wide association study reveals plant loci controlling heritability of the rhizosphere microbiome. The ISME journal, 15(11), 3181–3194.
Gao, C., Montoya, L., Xu, L., Madera, M., Hollingsworth, J., Purdom, E., … & Taylor, J. W. (2020). Fungal community assembly in drought‐stressed sorghum shows stochasticity, selection, and universal ecological dynamics. Nature Communications, 11(1), 34.
Grieneisen, L., Dasari, M., Gould, T. J., Björk, J. R., Grenier, J. C., Yotova, V., … & Blekhman, R. (2021). Gut microbiome heritability is nearly universal but environmentally contingent. Science, 373(6551), 181–186.
Kell, D. B. (2012). Large‐scale sequestration of atmospheric carbon via plant roots in natural and agricultural ecosystems: why and how. Philosophical Transactions of the Royal Society B: Biological Sciences, 367(1595), 1589–1597.
Panchal, P., Preece, C., Peñuelas, J., & Giri J. (2022). Soil carbon sequestration by root exudates. Trends in Plant Science, 27(8), 749–757.
Pascale, A., Proietti, S., Pantelides, I. S., & Stringlis, I. A. (2020). Modulation of the root microbiome by plant molecules: the basis for targeted disease suppression and plant growth promotion. Frontiers in Plant Science, 10, 1741.
Wagner, M. R. (2021). Prioritizing host phenotype to understand microbiome heritability in plants. New Phytologist, 232(2), 502–509.
Zhang, J., Liu, Y.X., Zhang, N., Hu, B., Jin, T., Xu, H., … & Bai, Y. (2019). NRT1. 1B is associated with root microbiota composition and nitrogen use in field‐grown rice. Nature Biotechnology, 37(6), 676–684.
Text © . The authors. CC BY-NC-ND 4.0. Except where otherwise noted, images are subject to copyright. Any reuse without express permission from the copyright owner is prohibited.